Scenarios
Early Feasibility Assessment (EFA) is the application of mechanistic PKPD models, built from first principles, and parameterized by data that is readily available early in drug discovery to make effective dose predictions.
EFA applies mechanistic PKPD models without fitting to PK and PD data, thus can be applied at an early stage even before the generation of the candidate or tool molecules. It leverages the ability of mechanistic models to utilize biological data from the literature that describes the intended pharmacology, the biophysical properties of the target, and physiological parameters such as compartment volumes, cell number, receptor expression levels, and soluble protein concentrations.
A "scenario" is the core object of EFA. It combines a model, parameters that model initial conditions and reaction rates, and a criteria of pharmacodynamic or pharmacological feasibility.
Model Parameters
Model parameters define the initial conditions and reaction rates of the model. They describe drug properties, such as binding affinity, and the rate of drug clearance, as well as target properties such as receptor expression, ligand concentration, and turnover.
Together, the drug and target parameters define pharmacokinetic and pharmacodynamic behavior of the model.
Metrics
For a drug to have biological activity, it must have some minimum pharmacokinetic or pharmacodynamic effect. EFA identifies drug properties necessary to feasibly achieve such an effect. The appropriate choice of a criteria by which to evaluate the feasibility of a drug and target depends on the specific biology of a disease indication and the target's role in it. In Applied BioMath Assess, these are called "Metrics".
For example, "sustained inhibition of TNF and TNFR binding relative to pre-treatment is greater than 90%" is a potential Metric.
The Metrics are specific to each model within Assess. For details on each criteria, see the model documentation inside Assess.
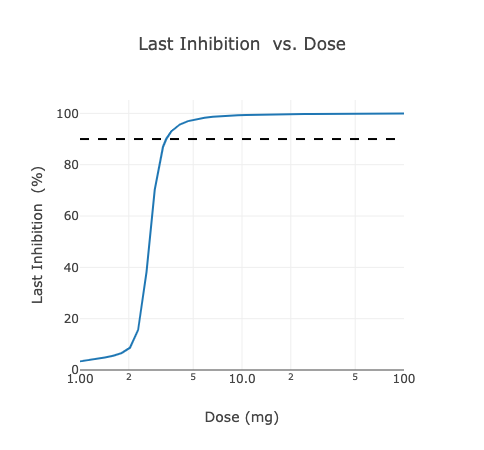
1D and 2D Scans
Applied BioMath Assess supports 1D and 2D scans. A scan varies 1 or 2 model parameters over a range to identify how the value of a metric changes. For instance, for an anti-receptor monoclonal antibody model, we can vary the dose of the therapeutic between 1 and 100 mg to identify the critical value at which 90% target inhibition occurs.